SCAN: Schistosomiasis Collection at the Natural History Museum
Natural History Museum's Aidan Emery outlines how the country's largest schistosomiasis collection is helping researchers to develop the tools and techniques to control and eliminate the parasites.
At the back of the whale hall in the Natural History Museum, down a flight of stairs, you will the Molecular Collections facility, home of SCAN, the schistosomiasis collection at NHM.
Part funded by the Wellcome Trust, SCAN is building a collection of the parasites that cause schistosomiasis, along with their snail intermediate hosts. Schistosome parasites are difficult to collect, so we want to facilitate research into this disease by providing genetically diverse specimens for the global research community.
In our busy office you might meet my three colleagues on the SCAN project, Fiona Allan, Muriel Rabone and Toby Landeryou.
Fiona, the SCAN manager, coordinates much of our fieldwork and collaboration. It’s much more efficient for us to work in partnership than alone. For example in the last few years we have worked alongside SCORE and ZELS with London Centre for NTD Research partners at NHM, Royal Veterinary College and Imperial College London and on collaborative projects in Niger, Senegal, Tanzania and Zanzibar. We can’t expect our project partners submit specimens to SCAN without providing any immediate benefits, so we get involved in collecting and sample handling to the extent that we are usually indistinguishable from the staff working on the partner project.

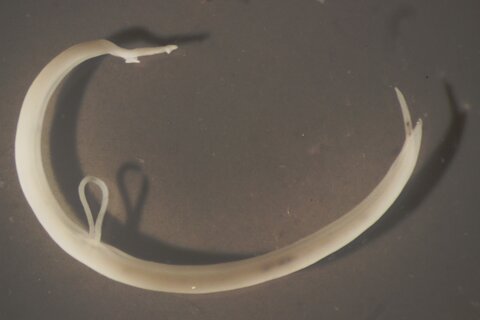
The adult schistosome worms that infect humans are not accessible for direct sampling because they live in the small veins around the intestine or bladder. However, eggs they produce migrate through the gut or bladder wall into urine or stool as a route out of the body. Unfortunately, many of the eggs don’t make it, and are swept along in the bloodstream until caught in the liver’s capillary network where they cause inflammatory responses responsible for much of the pathology associated with schistosomiasis. To collect schistosomes, we filter the eggs from stool or urine samples through a mesh, adding clean water to encourage them to hatch into miracidia, the short-lived, non-feeding life stage that infects water snails. We pipette each miracidium individually onto a DNA storage card that allows us to store the sample dry, at room temperature for future analysis.
Collecting schistosomes would be of little value without also assembling the data associated with the samples, and this is where our sample and data coordinator, Muriel Rabone comes in. Our partner projects have many different priorities and formats for collecting data and it is Muriel’s job to check the data, follow up and correct errors, and enter into our own spreadsheets and databases. By itself, this process is of enormous immediate advantage to studies analysing thousands of individual specimens from multiple locations. The data are uploaded into NHM’s collection management database, and made available through the SCAN project website and NHM’s data portal.
Opposite the office is the molecular collections facility itself, a purpose-built storage and preparation lab for molecular-grade specimens. We store our collections in several different formats, depending on sample type and how it was originally collected. The ultimate for preservation is liquid nitrogen, which we use to store our collection of adult schistosome parasites. These are the legacy of NHM collecting schistosomes for laboratory culture from the 1980s until around 2010. At the moment we are storing extracted DNA in -80°C freezers, and keeping DNA storage cards at ambient temperature. We also maintain the host snails, refrigerated, in molecular-grade ethanol.

Our newest collaboration is an exciting partnership with the Schistosomiasis Control Initiative (SCI) at Imperial College London. This also brings in our newest colleague, Toby Landeryou, who is in a joint SCAN/SCI post aiming to introduce routine genetic sampling into SCI’s survey procedures. SCI works primarily with national control programmes in sub-Saharan Africa. Including genetic sampling will enable the creation a reference bank in case of complications resulting from genetic changes in parasite populations such as drug resistance. We hope to undertake our first pilot collection within one of SCI’s surveys in the next few weeks.
Schistosomiasis is a major problem in parts of sub-Saharan Africa, is caused by several species of parasites, and transmitted by a number of different water snails. Other schistosome species infect livestock, with significant veterinary impact. Environmental change can alter which species of snails and parasite are found, and recent studies have shown that human and veterinary species of schistosome can hybridise, with unknown implications for control and epidemiology. This dynamic picture is why it is important to make the most of any sampling opportunities so we have a record of what is happening to the parasite and have references available for comparison.
Further resources
- Project - SCAN: Schistosomiasis Collections at the Natural History Museum
- Project - WHO Collaborating Centre for Identification & Characterization of Schistosome Strains & their Snail Intermediate Hosts
- Research article - Whole genome amplification and exome sequencing of archived schistosome miracidia
- Research article - Interrupting seasonal transmission of Schistosoma haematobium and control of soil-transmitted helminthiasis in northern and central Côte d’Ivoire: a SCORE study protocol
- Research article - Development of novel multiplex microsatellite polymerase chain reactions enabling high-throughput population genetic studies of Schistosoma haematobium.
- Research article - Whole genome resequencing of the human parasite Schistosoma mansoni reveals population history and effects of selection.